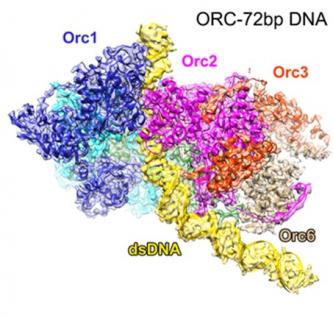
Cells propagate by making copies of themselves through replicating their DNA genome, which are blueprints of their identities. Every full grown human came from a single fertilized egg cell whose genome is replicated approximately 10 million-billion times. What does the molecular machine that carries out this Herculean task look like? A research team led by scientists from the Hong Kong University of Science and Technology (HKUST) have determined the three dimensional structure of the DNA replication machinery at atomic resolution for the first time in history.
When DNA replication was first proposed based on its double helix structure over half a century ago, many believed that deciphering the machine that separates the two strands of DNA for replication is near to come. However, it turns out to be a much complicated task due to the large size, multi-partite nature (made up of three engines) and its flexibility of the machine.
Today, capitalizing on the Cryo-electron microscopy (CryoEM) technology, a team led by Prof Bik Tye, Senior Visiting Member of HKUST Jockey Club Institute for Advanced Study (IAS) (retired Visiting Professor of Life Science (LIFS)) at HKUST and Prof Yuanliang Zhai, former Research Assistant Professor (RAP) at HKUST who is now an Assistant Professor at the University of Hong Kong, in collaboration with Prof Ning Gao, Professor of Life Sciences at Peking University – have managed to decipher the function of eukaryotes’ DNA replication machine, Origin Recognition Complex (ORC), at unprecedented resolution. This structure explains how ORC is able to scan a sea of bases (DNA is made up of 4 bases, A, T, G, C) to select the correct sites programmed for DNA replication to begin.
It is believed that indiscriminate selection of too many sites may lead to rapid replication of the genome and therefore rapid cell divisions – a characteristic of cancer cells. In contrast, inefficient selection of sites resulting in sluggish cell divisions especially at critical junctures of human development, such as embryogenesis, may lead to developmental disorders. A three dimensional view of the DNA replication machine at 3Å resolution may help to identify better targets for cancer therapy such that synthetic chemicals can be custom made to fit the target. More importantly, structures help to fully understand the mechanistic functions of molecular machines and therefore the roots of diseases due to suboptimal functions of these machines.
These findings on ORC were published in the prestigious scientific journal Nature on 4 July 2018 - the latest of a series of articles published by the Tye (HKUST)/Gao (PKU) collaboration that has opened the door for deciphering the function of the DNA replication machine at unprecedented resolutions. The first, published in Nature 2015, determined the structure of the core engine of the DNA replication machine called the MCM complex. The second reported an open-ringed structure of the Cdt1–Mcm2–7 complex as a precursor of the MCM double hexamer, which was published in Nature Structural and Molecular Biology.
Prof Tye’s interest in the mechanisms for DNA replication dated from when she established her own laboratory at Cornell University as an Assistant Professor. Her group published the initial paper in 1984 that identified the MCM2-7 genes as key components in DNA replication. Dr Yuanliang Zhai, formerly worked in Prof Tye’s group as RAP at HKUST’s LIFS and a Junior Fellow of IAS is now Assistant Professor in School of Biological Sciences at the University of Hong Kong. The co-authors of this Nature paper also include Dr Wai Hei Lam, Post-doctoral fellow at HKUST’s LIFS and Dr Yongqian Zhao, RAP at HKUST’s LIFS and Junior Fellow of IAS.
About The Hong Kong University of Science and Technology
The Hong Kong University of Science and Technology (HKUST) (www.ust.hk) is a world-class research university that focuses on science, technology and business as well as humanities and social science. HKUST offers an international campus, and a holistic and interdisciplinary pedagogy to nurture well-rounded graduates with global vision, a strong entrepreneurial spirit and innovative thinking. HKUST attained the highest proportion of internationally excellent research work in the Research Assessment Exercise 2014 of Hong Kong’s University Grants Committee, and is ranked as the world’s best young university in Times Higher Education’s Young University Rankings 2018. Its graduates were ranked 12th worldwide and top in Greater China in Global Employability University Survey 2017.
For media enquiries, please contact:
Anita Lam
Tel : 2358 6313
Email: anitalam@ust.hk
Johnny Tam
Tel : 2358 8556
Email : johnnytam@ust.hk